This notebook prepares a small, reproducible Zarr subset that is fast to iterate on for regridding experiments.
It opens the dataset from a Kerchunk catalog generated on HPC, and the same workflow can open:
directly on the HPC filesystem, or
over HTTPS via the DATARMOR export service (
https://data-fair2adapt.ifremer.fr/).
Goal¶
Produce a temporary Zarr (e.g. small.zarr) that is then used as input to:
regrid_apply_ufunc.ipynb
Outputs¶
OUT_ZARR: a lightweight subset written to Zarr (local or HPC scratch)
Steps¶
Open the dataset from a Kerchunk catalog (portable: HPC filesystem or HTTPS export)
Select variables and ensure lon/lat are explicit coordinates
Build a polygon mask helper for the model grid
Define the ROI and apply the mask (load ROI polygon, mask, and subset)
Write a temporary Zarr (e.g.
small.zarr) for use byregrid_apply_ufunc.ipynb
Tip: keep this subset small (few timesteps / ROI mask) until the workflow is validated.
Kerchunk catalog options¶
A Kerchunk catalog is a JSON mapping that lets Xarray open a collection of NetCDF (or similar) files as a virtual Zarr dataset.
Depending on where you run this notebook, you can point to:
HPC filesystem path (fast, when you have direct access to
/scale/...)HTTPS export (portable, when you access the same data through
https://data-fair2adapt.ifremer.fr/)
Below are example catalog paths that have been created previously (kept here as a reference).
1. Open the dataset from a Kerchunk catalog (portable: HPC filesystem or HTTPS export)¶
Many Kerchunk catalogs store references to the original files (often as absolute HPC paths).
When opening through HTTPS, we rewrite those references from:
/scale/project/lops-oh-fair2adapt/... → https://data-fair2adapt.ifremer.fr/...
The cell below:
Detects whether the HPC path exists (so we are running on the cluster).
Otherwise loads the JSON over HTTPS, patches references in-memory, and opens the dataset.
(Optional) can cache the patched references locally as a parquet file for faster re-opening.
%%time
import json
from pathlib import Path
import fsspec
import xarray as xr
from healpix_regrid import patch_kc_refs_inplace
HPC_PREFIX = "/scale/project/lops-oh-fair2adapt/"
HTTPS_PREFIX = "https://data-fair2adapt.ifremer.fr/"
CATALOG_PATH = "fpaul/tmp/riomar_3months.json"
OUT_PARQUET = "riomar_3months_.parq" # local parquet refs cache
# ------------------------------
# 1) HPC mode: open directly
# ------------------------------
if Path(HPC_PREFIX).exists():
KERCHUNK_CATALOG = HPC_PREFIX + CATALOG_PATH
print("Running in HPC mode:", KERCHUNK_CATALOG)
ds = xr.open_dataset(KERCHUNK_CATALOG, engine="kerchunk", chunks={})
# ------------------------------
# 2) HTTPS mode: fetch JSON, patch refs, open
# ------------------------------
else:
KERCHUNK_CATALOG = HTTPS_PREFIX + CATALOG_PATH
print("Running in HTTPS mode:", KERCHUNK_CATALOG)
with fsspec.open(KERCHUNK_CATALOG, "rt") as f:
kc = json.load(f)
kc = patch_kc_refs_inplace(kc)
ds = xr.open_dataset(kc, engine="kerchunk", chunks={})
ds2. Select variables and ensure lon/lat are explicit coordinates¶
The original dataset contains many variables. For this demo we keep:
temp(temperature)salt(salinity)zeta(sea surface height)
We also load the 2D longitude/latitude fields and attach them as coordinates (nav_lon_rho, nav_lat_rho).
Loading them explicitly avoids repeated remote reads later (plots, masking, regridding, etc.).
ds = ds[["temp", "salt", "zeta"]].assign_coords(
nav_lon_rho=ds["nav_lon_rho"].load(),
nav_lat_rho=ds["nav_lat_rho"].load(),
)
ds3. Build a polygon mask helper for the model grid¶
To extract a spatial subset, we load a boundary polygon (GeoJSON) and create a boolean mask on the dataset grid:
True→ grid point is inside the polygonFalse→ outside
This is useful to reduce the dataset to a Region Of Interest (ROI) before saving or regridding.
import numpy as np
from healpix_regrid import apply_polygon_mask4. Define the ROI and apply the mask (load ROI polygon, mask, and subset)¶
For the regridding demo we focus on a limited area defined by an outer boundary polygon (stored in outer_boundary.geojson).
We will:
Read the polygon from GeoJSON.
Trim the dataset in time (keep only a couple of timesteps for a lightweight example).
Add the polygon mask to the dataset.
In the next notebook (e.g. simple_regrid.ipynb) we will use the saved subset for regridding experiments.
import geopandas as gpd
# 1. Read the polygon from GeoJSON.
gdf = gpd.read_file("outer_boundary.geojson", driver="GeoJSON")
poly = gdf.geometry.iloc[0]
# 2. Trim the dataset in time
# (keep only a couple of timesteps for a lightweight example).
ds = ds.isel(time_counter=slice(0, 2))[["temp", "salt", "zeta"]]
# 3. Add the polygon mask to the dataset.
ds_temp = apply_polygon_mask(ds, poly)
ds_tempInspect the native grid extent for plotting
After applying the ROI mask, we compute the min/max longitude/latitude of valid grid points.
This helps confirm we are working on the expected geographic area and can be used to set plot limits.
lat = ds_temp["nav_lat_rho"]
lon = ds_temp["nav_lon_rho"]
valid = (lat != -1) & (lon != -1) # or (lat > -90) & (lon > -180) if you prefer
lat_min = lat.where(valid).min().item()
lat_max = lat.where(valid).max().item()
lon_min = lon.where(valid).min().item()
lon_max = lon.where(valid).max().item()
lat_min, lat_max, lon_min, lon_max(46.89809036254883, 47.4329719543457, -2.8933334350585938, -1.90666663646698)Quick visual check
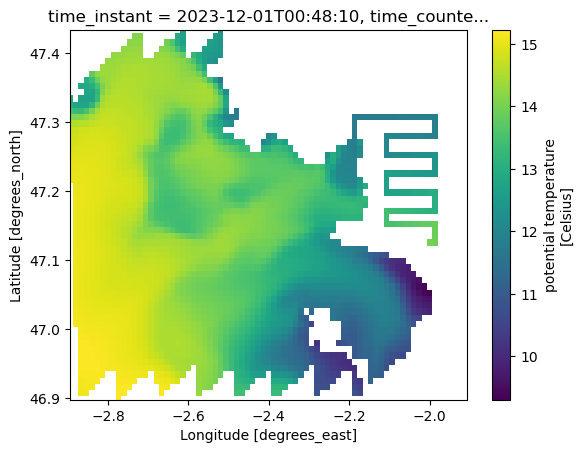
Plot one surface slice (time_counter=0, s_rho=0) to verify that:
coordinates are correctly attached (
nav_lon_rho,nav_lat_rho)the selected subset matches the intended area
ds_temp.temp.isel(time_counter=0, s_rho=0).plot(
y="nav_lat_rho", x="nav_lon_rho", ylim=(lat_min, lat_max), xlim=(lon_min, lon_max)
)
5. Write a temporary Zarr (e.g. small.zarr) for use by regrid_apply_ufunc.ipynb¶
To make the next steps (regridding) fast and reproducible, we write the masked / subsetted dataset to a local Zarr store.
Tip: adjust OUT_ZARR to a location that exists on your machine (or to a shared scratch directory on HPC).
%%time
OUT_ZARR = "/Users/todaka/data/RIOMAR/small.zarr"
ds_temp.to_zarr(OUT_ZARR, mode="w")/Users/todaka/micromamba/envs/pangeo/lib/python3.13/site-packages/zarr/api/asynchronous.py:247: ZarrUserWarning: Consolidated metadata is currently not part in the Zarr format 3 specification. It may not be supported by other zarr implementations and may change in the future.
warnings.warn(
CPU times: user 12.7 s, sys: 7.4 s, total: 20.1 s
Wall time: 1min 56s
<xarray.backends.zarr.ZarrStore at 0x13ff727a0>Re-open the Zarr subset and plot
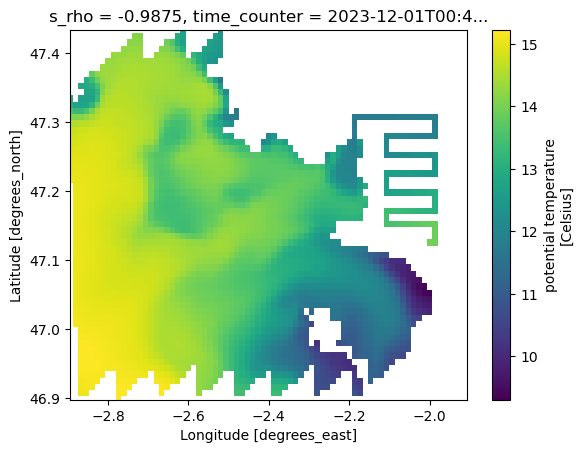
This verifies that the data were written correctly and can be opened independently of the original Kerchunk catalog.
%%time
OUT_ZARR = "/Users/todaka/data/RIOMAR/small.zarr"
small_ds = xr.open_zarr(OUT_ZARR)
small_ds.temp.isel(time_counter=0, s_rho=0).plot(
y="nav_lat_rho", x="nav_lon_rho", ylim=(lat_min, lat_max), xlim=(lon_min, lon_max)
)CPU times: user 40.3 ms, sys: 8.61 ms, total: 48.9 ms
Wall time: 48.4 ms
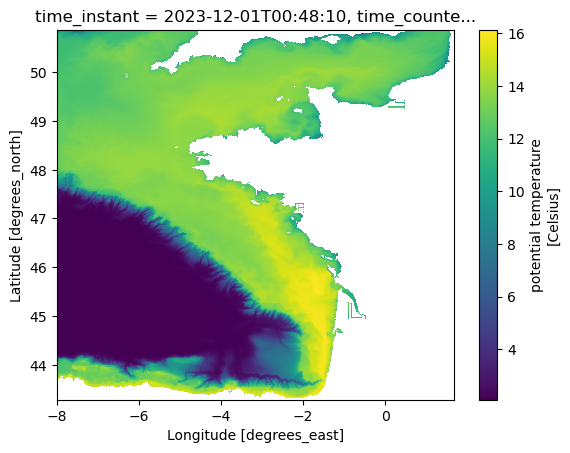
Optional: broader view
A second plot with wider latitude limits can help spot any unexpected artifacts (e.g., missing values, wrap-around).
lat = ds["nav_lat_rho"]
lon = ds["nav_lon_rho"]
valid = (lat != -1) & (lon != -1) # or (lat > -90) & (lon > -180) if you prefer
lat_min = lat.where(valid).min().item()
lat_max = lat.where(valid).max().item()
lon_min = lon.where(valid).min().item()
lon_max = lon.where(valid).max().item()
ds.temp.isel(time_counter=0, s_rho=0).plot(
y="nav_lat_rho", x="nav_lon_rho", ylim=(lat_min, lat_max), xlim=(lon_min, lon_max)
)
(Optional) Convert ROI data to an XDGGs / HEALPix-indexed DataArray
If your next step is to move from the native model grid to a DGGS representation (e.g. HEALPix),
it can be convenient to store values against a cell_ids index.
The helper below builds an xarray.DataArray that follows the metadata conventions expected by xdggs.
You can then attach this to a dataset, write it to Zarr, or use it in downstream regridding/resampling code.
import xarray as xr
import xdggs
def make_xdggs_dataarray_from_cell_ids(
data,
cell_ids,
level: int = 15,
name: str = "da",
):
"""
Build an xdggs-compatible DataArray indexed by HEALPix `cell_ids`.
Parameters
----------
data : array-like
Values for each cell (same length as `cell_ids`).
cell_ids : array-like of int
HEALPix cell identifiers (nested indexing).
level : int
HEALPix resolution level.
name : str
Name of the DataArray.
Returns
-------
xr.DataArray
Decoded and enriched with latitude/longitude coordinates via xdggs.
"""
cell_ids = np.asarray(cell_ids, dtype=np.int64)
da = xr.DataArray(
data,
dims=("cell_ids",),
coords={"cell_ids": ("cell_ids", cell_ids)},
name=name,
)
# Minimal metadata expected by xdggs
da["cell_ids"].attrs.update(
{
"grid_name": "healpix",
"level": int(level),
"indexing_scheme": "nested",
"ellipsoid": "WGS84",
}
)
# Decode -> attach DGGS accessor -> compute lat/lon coordinates
return da.pipe(xdggs.decode).dggs.assign_latlon_coords()
with np.load("parent_ids.npz") as data:
parent_ids = data["parent_ids"]
parent_level = int(data["parent_level"])
da_hp = make_xdggs_dataarray_from_cell_ids(
np.ones(len(parent_ids)), parent_ids, level=parent_level
)
da_hp.dggs.explore(alpha=0.3)